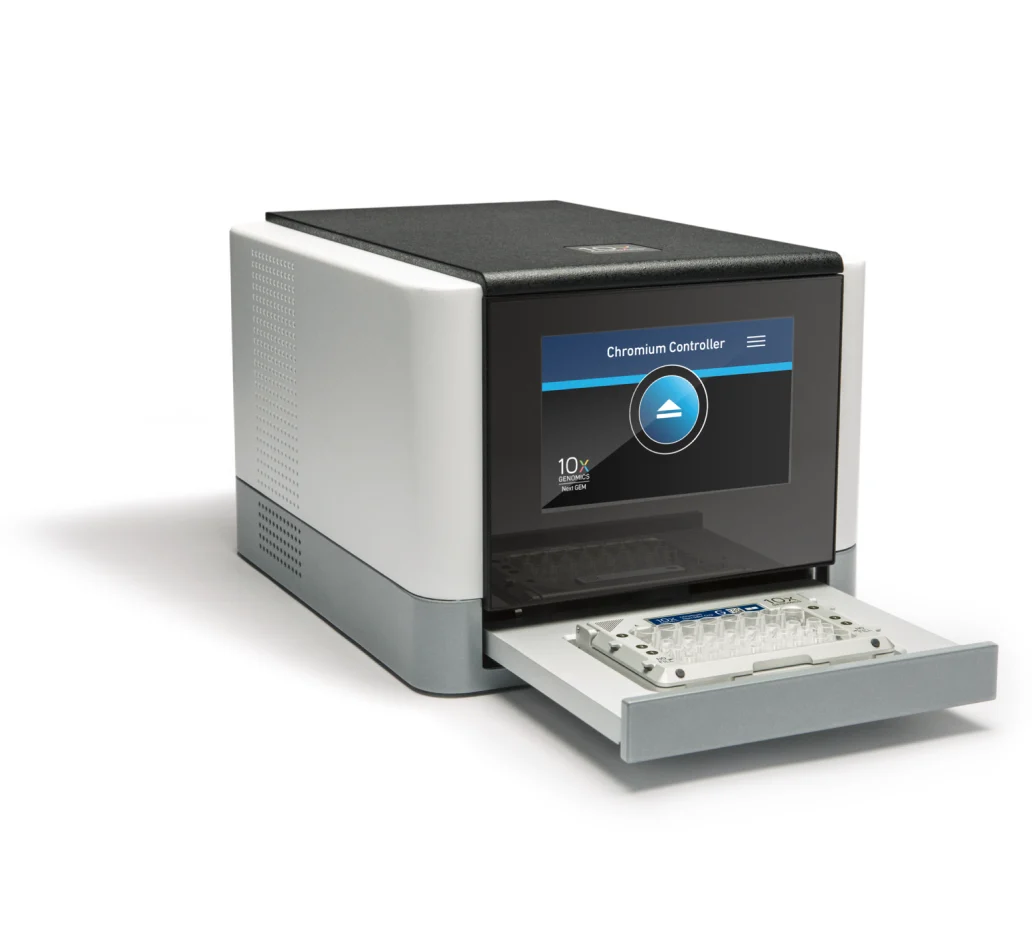
  
Detection of multiplets and their removal prior to analysisĪs part of their TotalSeq antibody catalog, BioLegend offers this technology through antibody-oligonucleotide conjugates that can be used for both CITE-seq and Cell Hashing.Increased number of possible replicates in a single experiment.Increased number of cells assayed in a single experiment.Increased sample throughput in a single experiment.Cells assigned a given single tag are binned together, bioinformatically recapitulating the individual samples originally mixed together. Tag assignment enables identification of droplets that originally contained one (singlet) or more cells (multiplets). After cell encapsulation, library preparation, and sequencing, molecular tag information is assigned to cells.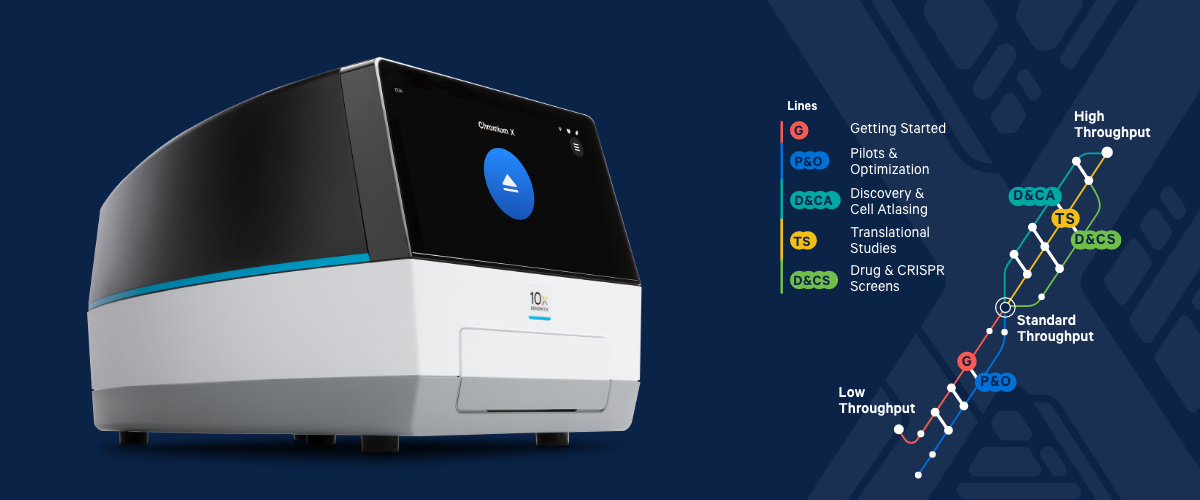 
The set of multiplexed samples are processed together in a single GEM well. Mostly this comes in two forms – looking for cells that are expressing certain proteins on the cell surface (CITE-seq) or using ubiquitous markers in order to pool different samples together into one channel of the chip (Cell Multiplexing/Cell Hashing).Ĭell or nuclei samples can be labeled with a molecular tag and subsequently mixed with other labeled samples. Single-cell data are analyzed with the Cell Ranger and Loupe Cell Browser software.įeature Barcoding is the broad term used by 10x Genomics in order to describe any method that adds extra layers of information to cells by running single cell gene expression in parallel with other assays to gain useful biological information. Quenching Buffer + Enhancer + 50% Glycerol from 10X Fixation kitġX Nuclei Buffer with RNase Inhibitor and DTT Other standard media with no more than 10% FBS or 2% BSA No EDTA Standard: 10,000 cells per channel up to 80,000 cells per chip Multiplex: 128,000 cells per channel up to 1,024,000 cells per chip Singleplex: 10,000 cells per channel up to 80,000 cells per chip 
High: 20,000 cells per channel up to 320,000 cells per chip up to 730,000 cells per chip with plexing Standard: 10,000 cells per channel up to 80,000 cells per chip 140,000 cells per chip with plexing High: 20,000 cells per channel up to 320,000Ĭells per chip up to 730,000 cells per chip with plexing Standard: 10,000 cells per channel up toĨ0,000 cells per chip 140,000 cells per chip Simultaneous detection of mRNA and chromatin accessibilityįixed cells, fixed nuclei, fixed tissue, FFPE tissue Probe-based whole transcriptome gene expression Please see our CSCI Flow Cytometry 10x Genomics Service document for more pricing and more details.Flavors of 10x Genomics Single Cell Assays Users must complete comprehensive training with CSCI Flow Cytometry staff before being granted access to the Chromium Control and workspace.Researchers whose needs are infrequent are encouraged to utilize the SCC's full-service option. The service is recommended for researchers who require off-hours access to the Chromium Controller and plan to use the technology regularly. 10x Genomics access is offered on a self-use basis only.Users can obtain 10x Next GEM reagents from the SCC when using the core's library preparation and sequencing services.Library preparation and sequencing must be arranged with the SCC or independently by researchers. CSCI Flow Cytometry supports only the GEM-capture and cDNA barcoding steps.Users also have the option to perform library preparation and arrange for sequencing independently. Barcoded cDNA can then be taken to the SCC for library preparation or sequencing. The Columbia Stem Cell Initiative Flow Cytometry core, in collaboration with the Columbia Genome Center Single CellĪnalysis Core (SCC) is pleased to offer access to 10x Genomics single-cell technology.Īfter obtaining reagents from the Single Cell Analysis core, fully-trained users have independent access to the 10x Genomics Chromium Controller, workspace, and associated equipment for the generation of barcoded cDNA from GEM-captured cells. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
